Maize Genetics Cooperation Newsletter vol 88 2014
RIO CUARTO, ARGENTINA
Universidad Nacional de RÔøΩo Cuarto
Stalk and root rot in maize hybrids: analysis
of gentoype ×
environment interaction using GGE biplots
Fissore, MJ; IbaÔøΩez, MA; Bonamico, NC AND Di Renzo,
MA.
Mejoramiento GenÔøΩtico,
Facultad de AgronomÔøΩa y Veterinaria, Universidad Nacional de RÔøΩo Cuarto,
Agencia NÔøΩ 3, 5800 RÔøΩo Cuarto, Argentina. e-mail: [email protected]
INTRODUCTION
Stalk and root rot (SRR) is one of the most important maize (Zea mays L.) crop diseases in Argentina, as
it is widely spread and has high incidence and severity (Buchaca,
2008). The most efficient, economical and environmentally sustainable control
method involves the use of resistant genotypes. The disease presents diverse
symptoms with different degrees of manifestations and incidence values among
the analyzed populations. As a consequence, resistance indices have been
proposed (Grau et al., 1982); here we propose
the multidimensional indicator of stalk and root rot (MISRR). The response of
the different hybrid genotypes to the disease is influenced by environmental
conditions; therefore, evaluating the materials in different environments is an
interesting approach. The differential genotype response to a disease in diverse
environments results in a genotype ×
environment interaction; quantifying the magnitude of the interaction is of
interest when evaluating genotypes.
To
describe genotype adaptability and stability, different statistical methods that explore the genotype × environment interaction and estimate it
with improved accuracy have been proposed. Although the phenotypic character of
interest is the result of the effects of genotype (G), environment (E) and their interaction (G×E), only the main effects of G and of G×E are relevant to genotype evaluation (Yan
et al., 2000; Yan
and Tinker, 2006). Unlike the
analysis of variance (ANOVA) and comparison of means, which only allow us to
select those genotypes that are above a given yield level, sites regression
models (SREG) (Cornelius et al., 1996; Crossa and
Cornelius, 1997; Crossa et al., 2002) are
recommended when environment effects are an important source of variation. These
models have the additional advantage of providing a graphical analysis known as
GGE biplot (Yan et al., 2000; 2001). The aim
of the present work was to study G×E
interaction via a multidimensional indicator of SRR in maize using the GGE biplot analysis.
MATERIALS AND METHODS
The multidimensional indicator of stalk and
root rot (MISRR) was estimated in 12 maize hybrids from different seed
companies (Table 1) based on field evaluations conducted in three localities from
the semiarid region of Argentina during a crop season: Buchardo
(34ÔøΩ45`19``S, 63ÔøΩ31`27``W) and Olaeta (CÔøΩrdoba
province; 32ÔøΩ58`52``S, 63ÔøΩ47`50``W) and Papagayo (San
Luis province; 32ÔøΩ42`41``S, 65ÔøΩ11`47``W). The
hybrids were selected because they are the most widely used by the AsociaciÔøΩn Argentina de Consorcios
Regionales de ExperimentaciÔøΩn
AgrÔøΩcola (AACREA: Argentine Association of Regional Consortiums
for Agricultural Experimentation).
Table
1. Seed companies and name of the evaluated commercial hybrids
|
Seed companies |
Hybrid |
Seed companies |
Hybrid |
|
Illinois |
I880MG |
Pannar |
PANPEX168MG |
|
KWS ARG |
KM4911TDMAX |
Pionner |
P1979Y |
|
La Tijereta |
LT622MGRR2 |
Pionner |
P2053Y |
|
La Tijereta |
LT632MG |
Pionner |
P2069Y |
|
Monsanto |
DK190MGRR2 |
Syngenta |
NK880TDMAX |
|
Monsanto |
DK747MGRR2 |
Syngenta |
NK910TDMAX |
The indicator (MISRR) is a measure of the response
to the disease and is classified according to the simultaneous expression of
the selected symptoms, weighted by the incidence of the different degrees along
a scale in a set of plants of each hybrid. MISRR can range from 0 to 100, with
100 corresponding to severely affected genotypes.
![]()
The following rating scale was
defined: 0= no symptoms, 1= change in pith color (dark or pink) or presence of
stripes on the basal internodes; 2= pith disintegration or changes in color and
striping in the lower internodes, 3= pith disintegration, with change in color
and presence of stripes.
The experimental design used in the three localities was a complete random
block, with three replications. Each sample consisted of five plants; the
number of plants with each one of the symptoms of stalk and root rot (infected)
and the number of healthy plants were counted to determine the incidence of
each symptom.
The data obtained across
environments was analyzed using ANOVA and SREG statistical model for
multi-environments (Cornelius et al., 1996; Crossa
and Cornelius, 1997; Crossa et al., 2002), which
includes G + G×E in the bi-linear
term. This model provides an easy-to-understand graph known as GGE biplot (Yan et al., 2000; 2001), which facilitates
the analysis of genotype behavior as well as the effect of the interaction with
the environment. This graph is built based on the first two principal
components (PCs) of the SREG model. The use of the biplot allows
us to identify the genotypes of highest production in each environment as well
as those of highest stability. In
addition, the model shows clustering of genotypes and environments with similar
response pattern and identifies the environments that are most representative and
that can best discriminate among genotypes. Experimental data were analyzed
with Infogen statistical software (Balzarini et al.,
2013).
RESULTS
Results of the ANOVA of the MISRR for the 12 hybrids
evaluated in the three localities are shown in Table 2. The results evidence a
statistically significant G×E
interaction for the studied character (p<0.001); i.e., differences between means
of hybrids varied depending on the environments where they were evaluated.
Table 2. Mean square and
significance of the analysis of variance of the multidimensional indicator of
stalk and root rot for 12 maize commercial hybrids evaluated in Buchardo, Olaeta and Papagayo.
|
SV |
df |
MS |
p |
|
Locality (E) |
2 |
11224.84 |
0.0009 |
|
Hybrid (G) |
11 |
1501.99 |
0.0002 |
|
Locality>Block |
6 |
404.87 |
0.3941 |
|
Hybrid*Locality (G×E) |
22 |
1085.10 |
0.0006 |
|
Error |
66 |
381.13 |
|
SV: Source
of variation; df: degrees of freedom; MS: Mean square
The response of the maize
commercial hybrids was analyzed simultaneously for the three localities with
the biplot obtained from the SREG model (Figure 1). The
GGE biplot resulting from the analysis of the two
first PCs accounted for 90% of MISRR variability.
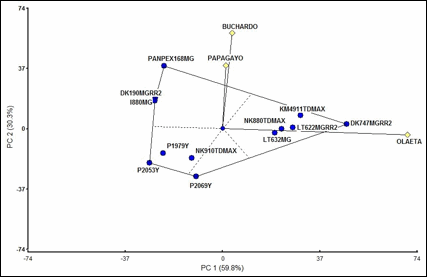
Figure 1. GGE biplot for the multidimensional indicator
of stalk and root rot (MISRR) of 12 maize commercial hybrids analyzed in the
localities of Buchardo, Olaeta
and Papagayo.
In the three
evaluated localities, environmental conditions influenced disease manifestation
in the different hybrids. The SREG model and graph analysis showed, on the one
hand, a group composed of the localities of Buchardo and
Papagayo, with hybrids exhibiting a similar response
pattern, with intermediate MISRR values, low discriminating ability and
representativeness of the tybrids. On the other hand,
the locality with greatest discriminating capacity and the most representative
one (the ideal environment) was Olaeta, where the
genotypes exhibited the greatest differences in reaction to the disease, therefore
allowing a better genotype characterization.
The values of the multidimensional
indicator analyzed via the biplot allowed us to identify
the response of the evaluated materials. The greatest interaction was detected
in the hybrid PANPEX168MG, a genotype specifically adapted to Buchardo and Papagayo
environments. Genotype DK747MGRR2 exhibited the poorest behavior (i.e., the
highest MISRR score) in all the environments. The hybrids P2053Y, I880MG and
DK190MGRR2 exhibited the best behavior and were the most stable ones, with low MISRR
values in all the evaluated environments (i.e., ideal genotype).
DISCUSSION
The GGE biplot was an efficient visual tool to reveal the interrelations
among G, E and G×E interaction. This
tool was used in studies focused on selection by multiple characters,
comparison of selection strategies and in the analysis of diallel
data to estimate combinatory ability (Yan et al., 2000; Yan and Tinker, 2006; Yan, 2013). The G×E interaction observed might be explained by elements that
compose the indicator. Indeed, as it is well known, genotypes can be resistant
to a given causal agent of stalk and root rot and be susceptible to others.
Thus, the environments with intermediate disease levels, such as those of the
group composed of Buchardo and Papagayo,
might have exhibited not only a high pathogen pressure but also a great
diversity of pathogens. Hence, all the materials analyzed were largely affected
by the disease, hindering the possibility of being differentiated. By contrast,
in Olaeta, the environment characterized by better
hydric conditions, only a few causal agents might have generated high pathogen
pressure. For this reason, certain genotypes susceptible to these microorganisms
were distinguished by higher indicator values, although they exhibited the
lowest values in the other two localities, therefore generating the G×E interaction. The greatest interaction
was represented by the hybrid PANPEX168MG, with low MISRR levels in Buchardo and Papagayo, and very
high ones in Olaeta. The genotype DK747MGRR2 is the
least recommendable one because it exhibited high susceptibility in all the environments.
Three hybrids, P2053Y, I880MG and DK190MGRR2, were found to be the most
recommendable ones for their tolerance and stability in all the environments. These
possible differences in the composition of microorganisms responsible for stalk
and root rot in each locality were not represented by MISRR. Hence, although
this indicator was a very useful tool to differentiate hybrids in Olaeta, in certain environments implementing MISRR may
require complementary studies for pathogen identification. Another possibility
is that the indicator may be used only when artificial inoculation is performed
with a single isolated microorganism, ensuring the uniform presence of the
pathogen in all genotypes, as proposed by Todd and Kommedahl
(1994), Buchaca (2008) and Szőke
et al. (2009).
The results
indicate that GGE biplot accounts for a high
proportion of the total variation of the data estimated for MISRR. Therefore,
the graph describes the interrelations between genotypes and environments, and
is used for the identification of hybrids and environments that show a good response
to the disease.
REFERENCES
Balzarini, M.G. y J.A. Di Rienzo. InfoGen
versiÔøΩn 2013. FCA, Universidad
Nacional de CÔøΩrdoba, Argentina. URL http://www.info-gen.com.ar
BUCHACA, M.B. 2008. Susceptibilidad a Fusarium verticillioides, Gibberella
zeae y Diplodia maydis en lÔøΩneas e hÔøΩbridos pre-comerciales
de maÔøΩces templados y tropicales. Tesis de EspecializaciÔøΩn en Mejoramiento
GenÔøΩtico Vegetal. Escuela para Graduados Alberto Soriano, Facultad de
AgronomÔøΩa, UBA, Buenos Aires, Argentina. 23 p.
CORNELIUS, P.L.; J. CROSSA and M.S. SEYEDSADR. 1996.
Statistical test and estimators of multiplicative models for
genotype-by-environment interaction. En: Genotype by environment interaction.
Boca Raton, FL, EE.UU. p: 199–234.
CROSSA, J. and P.L. CORNELIUS. 1997. Sites regresion and shifted multiplicative model clustering of
cultivar trials sites under heterogenety of variantes. Crop Science 37: 406–415.
CROSSA, J.; P.L. CORNELIUS and W. YAN. 2002. Biplots of linear-bilinear models for studying crossover
genotype × environment interaction.
Crop Sciencie 42: 619–633.
GRAU, C.R.; V.L. RADKE and F.L. GILLESPIE. 1982. Resistance of soybean
cultivars to Sclerotinia sclerotiorum. Plant Disease. 66: 506–508.
SZŐKE, C.; T. ÔøΩRNDÔøΩS; P. BÔøΩNIS y ÔøΩ.
SZÔøΩCSI. 2009. Fusarium stalk rot: a biotic stress
factor decisive for maize stalk strenght. VIII. Alps-Adria
Scientific Workshop. Neum, Bosnia-Herzegovina. p:
337–340.
TODD, L.R. and T. KOMMEDAHL. 1994. Image
analysis and visual estimates for evaluating disease reactions of corn to Fusarium stalk rot. Plant Disease 78: 876–878.
YAN, W. 2001. GGE biplot
Windows application for graphical analysis of multienvironment
trial data and other types of two-way data. Agronomy Journal 93: 1111–1118.
YAN, W. 2013. Biplot
analysis of incomplete two-way data. Crop Science 51: 48–57.
YAN, W. and N.A. TINKER. 2006. Biplot analysis of multi-environment trial data: Principles
and applications. Canadian Journal of Plant Science 86: 623–645.
YAN, W.; L.A. HUNT; Q. SHENG and Z. SZLAVNICS.
2000. Cultivar evaluation and mega-environment investigation based on the GGE biplot. Crop Science 40: 597–605.
Please Note: Notes submitted to the Maize Genetics Cooperation Newsletter may be cited only with consent of authors.