Maize Genetics Cooperation Newsletter vol 84 2010
Please Note: Notes submitted to the Maize Genetics
Cooperation Newsletter may be cited only with consent of authors.
IRKUTSK, RUSSIA
Institute of Plant physiology and Biochemistry,
Russian Academy of Sciences
NOVOSIBIRSK, RUSSIA
Institute of Cytology and Genetics, Russian
Academy of Sciences
Zea mays L. mitochondrial Mn-superoxide dismutase:
evidence in favour of potential DNA protective function
--Katyshev, AI; Subota, IY;
Deineko, EV; Konstantinov, YM
It is well known that a number of environmental stresses can lead to enhanced production of superoxide anion (O2-) radical within maize plant tissues, and plants are believed to rely on the enzyme superoxide dismutase (SOD) to detoxify this reactive oxygen species (ROS). SOD gene family in eukaryotes and prokaryotes consists of multiple genes encoding SOD isoforms. Plant cells contain almost all known SOD types which differ by their metal cofactor and subcellular location. Based on the metal cofactor used by the enzyme, SODs are classified into three groups: iron SOD (FeSOD), manganese SOD (MnSOD), and copper-zinc SOD (Cu/ZnSOD). FeSOD isoforms are located in the chloroplasts, MnSOD isoforms are in the mitochondria and the peroxisomes, and Cu-Zn SOD isoforms are in the chloroplasts, the cytosol, and the extracellular space. In this note we describe some biochemical properties of one MnSOD isoform in mitochondria. We suggest that this MnSOD isoform is involved in mitochondrial DNA protection from ROS.
Total RNA isolation from 3-day-old etiolated
hybrid VIR46MV seedlings was
performed by QIAGEN RNeasy Mini Kit according manufacturer�s instructions. cDNA
synthesis was carried out using the Promega Universal RiboClone cDNA Synthesis
System. Translated sequence of cDNA of
MnSOD3.1 gene flanked by NcoI and BglII sites was amplified using sequence-specific primers (5�-CGACCAAAGCCATGGCTCT-3�
as a forward and 5�-CCGTTAAGACAGATCTAGCAAGAACA-3� as a reverse primer). Primers for reverse transcription (RT)–PCR of the SOD3.1 gene were
designed using the VectorNTI5 program (Bethesda, USA). The sequence of MnSOD3.1
was obtained from GenBank database at http://www.ncbi.nlm.nih.gov/nucleotide/
(Acc. number M33119). Obtained
PCR-product was eluted from agarose gel using QIAEXII kit (QIAGEN, USA),
digested by NcoI and BglII enzymes and ligated into pQE60
expressing vector (QIAGEN, USA). Positive clones were analyzed by PCR and
subsequent sequencing on ABI PRISM 377 DNA automated sequencer as
recommended by Applied Biosystems.
The expression of recombinant protein was analyzed in total protein extracts in
combination with superoxide dismutase activity gel assay according Garnik et
al. (Russ. J. Plant Phys. 51:386-391, 2004). Elution of recombinant protein was
performed on Ni-NT agarose column (QIAGEN, USA) according manufacturer�s
instructions.
Analysis of DNA cleavage
activity of recombinant protein was performed as described by Chen et al. (Arch. Biochem. Biophys. 404:218-226, 2002) using pBlueScript KS II (+) plasmid DNA isolated by using GeneJet
Plasmid purification kit (Fermentas, Lithuania). In vitro
phosphorylation of MnSOD was conducted as described by Subota et al. (Russ. J. Plant Phys. 57:37-44, 2010).
Maize MnSOD3.1 gene
was cloned in bacterial expression vector pQE60 and recombinant protein was
isolated (Figure 1).
According to Fridovich (J. Biol. Chem. 272:25071-25076,
1997) and Descheneau, Newton (Int.
Congr. Plant Mitochondrial Biol. P.23, 2005) MnSOD in
bacteria and plants is DNA-associated protein which protects DNA against
oxidative damage by reactive oxygen species. We hypothesized that to perform this
function MnSOD should not possess DNA cleavage activity mediated through
generation of hydroxyl radical in Fenton reaction. To test this suggestion we
compared DNA cleavage activity of commercial bovine Cu-ZnSOD (Sigma, USA) and
recombinant maize MnSOD. Figure 2 shows that only in the case of 5 and 10 �g of bovine CuZnSOD cleavage
of plasmid DNA is observed (the quantity of linear form of plasmid DNA is
increased). So, maize MnSOD does not possess DNA cleavage activity in contrast
with Cu-ZnSOD.
It is known that the operation of mitochondrial
electron transport chain is one of the main sources of ROS in the cell.
Phosphorylation of MnSOD indicates that mitochondrial metabolism of ROS is presumably
regulated (Bykova et al., FEBS Lett. 540:141-146, 2003). Figure 3 shows that maize MnSOD enzyme is a phosphorylated protein.
Moreover, our analysis revealed that there are two phosphorylated MnSOD3.1
forms – native tetrameric and unusual dimeric form. Both of these forms
possess superoxide dismutase activity (Figure 3).
We suggest that some not known yet
properties of mitochondrial MnSOD may create background for special role of
this enzyme in mitochondrial DNA protection from reactive oxygen species. This
work was financially supported in part by the Integration projects SB RAS 7,
83, 98 and RFBR projects 08-04-01426, 09-04-00992.
Figure legends
Figure 1. Isolation of recombinant maize MnSOD3.1 protein. A - Electrophoretical analysis of bacterial total protein extracts. 1-5 –
different transformed bacterial clones. B
- Superoxide dismutase activity
analysis of recombinant proteins. 1, 2 – clones. C - Isolation of recombinant protein from bacteria. 1- lysate, 2
– flow-through, 3 – washing, 4 –eluate.
Figure 2. Analysis
of DNA cleavage activity of SOD. 1- plasmid DNA, 2-5 – plasmid
DNA + maize MnSOD (1, 2.5, 5 and 10 �g, respectively), 6-9
- plasmid DNA + bovine CuZnSOD (1, 2.5, 5 and 10 �g, respectively). C – coiled DNA, L –
linear DNA, SC – supercoiled DNA.
Figure 3. Phosphorylation of recombinant maize MnSOD3.1. 1 – protein
kinase-containing chromatography fraction of mitochondrial proteins, 2 – phosphorylation of
recombinant MnSOD3.1 by this fraction, 3 - superoxide dismutase activity analysis of phosphorylated MnSOD3.1. Narrows
indicate two forms of MnSOD3.1.
Figure 1.
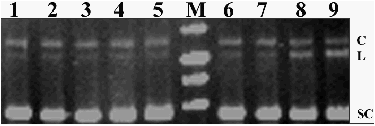
Figure 2.

Figure 3
